Meta-analysis is a quantitative method that synthesizes results from multiple independent studies to identify overall patterns, effect sizes, and sources of variability for specific treatments or research questions. It is particularly powerful for summarizing existing evidence and placing new findings within the context of current scientific trends through transparent and reproducible statistical integration of prior studies.
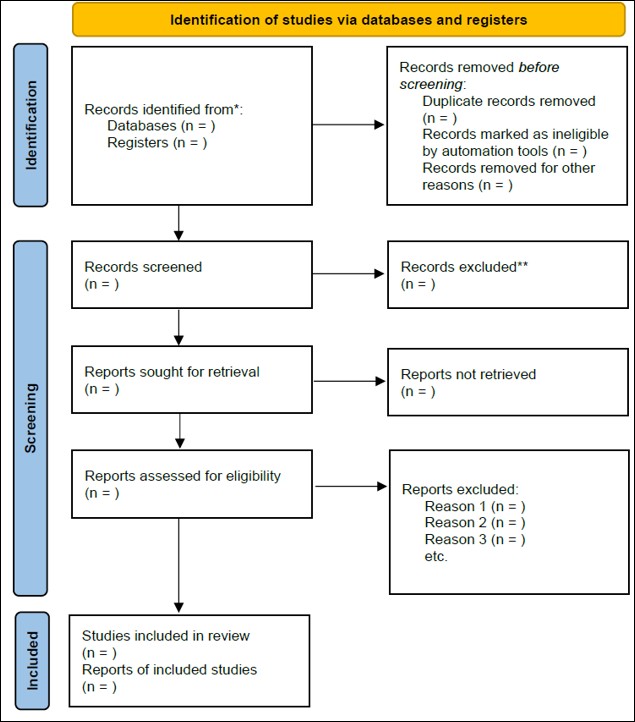
A rigorous meta-analysis begins with a comprehensive and systematic literature search using predefined search terms and Boolean combinations across multiple databases. Study selection and reporting are commonly guided by the PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) framework (https://www.prisma-statement.org), which requires explicit documentation of the identification, screening, eligibility, and inclusion of studies, as well as presentation of the search strategy and flow diagram.
Many studies labeled as “meta-analyses” fail to meet these standards. In such cases, study selection is often based on unsystematic or author-selected searches, leading to a biased evidence base. Without a transparent search strategy and adherence to PRISMA guidelines, these analyses cannot be considered true meta-analyses, as the results may reflect selection bias rather than the underlying research landscape.
Following established guidelines such as PRISMA is critical to ensure Transparency, Reproducibility, and Objectivity. Systematic search strategies and clearly defined inclusion criteria reduce selection bias and prevent authors from unintentionally—or intentionally—favoring studies that support a desired conclusion. Adherence to these rules allows readers and reviewers to assess the validity of the evidence base, replicate the analysis, and trust that the reported results represent the full scope of available research rather than a selectively curated subset.

To facilitate such searches for Meta-analysis, I recently developed a new R function, scopusmining(), for mining papers from the Scopus database.
1) Install the scopusmining() package
if(!require(remotes)) install.packages("remotes")
if (!requireNamespace("scopusmining", quietly= TRUE)) {
remotes::install_github("agronomy4future/scopusmining", force= TRUE)
}
library(remotes)
library(scopusmining)
2) Search strings
2.1) Enter a SCOPUS API key
Before running scopusmining(), it is required to enter a SCOPUS API key.
Sys.setenv(SCOPUS_API_KEY= "0000000000000000000000000000")
You can create a new SCOPUS API Key in the below website.
scopusmining()requires execution from an institutional IP address associated with an active Scopus subscription. API requests initiated from personal or non-institutional IP addresses may not be recognized by the Scopus API, resulting in failed queries despite the use of a valid API key.
2.2) “AND” logic
First, I search for the keywords Soybean AND Yield in the title, and Grain AND Pod in the abstract, restricting the publication years to 2020–2023. Therefore, I run the following code.
reference= scopusmining(
title_blocks= list(
list(terms= c("Soybean", "Yield"), logic= "AND")),
abskey_blocks= list(
list(terms= c("Grain", "Pod"), logic= "AND")),
years= 2020:2023,
dedupe= FALSE
)
The literature search was conducted using the scopusmining() function with the following search query:
TITLE(("Soybean" AND "Yield")) AND TITLE-ABS-KEY(("Grain" AND "Pod"))
AND DOCTYPE(AR) AND SRCTYPE(j)
This search strategy was documented and reported in the Meta-analysis to ensure transparency and reproducibility.
Let us check how many papers were retrieved. This search returned 104 papers.
nrow(reference$data) [1] 104
Let’s export this papers to .csv file.
write.csv(reference$data, "reference_list.csv", row.names= FALSE)
The reference list was exported as a .csv file.

2.3) “OR” logic
Next, I search for the keywords Soybean OR Yield in the title, and Grain OR Pod in the abstract, restricting the publication years to 2020–2023. I run the following code.
reference= scopusmining(
title_blocks= list(
list(terms= c("Soybean", "Yield"), logic= "OR")),
abskey_blocks= list(
list(terms= c("Grain", "Pod"), logic= "OR")),
years= 2020:2023,
dedupe= FALSE
)
The search query used in this search is presented as follows.
TITLE(("Soybean" OR "Yield")) AND TITLE-ABS-KEY(("Grain" OR "Pod")) AND DOCTYPE(AR) AND SRCTYPE(j)
This search returned 800 papers.
nrow(reference$data) [1] 800
This is expected because the search used the “OR” operator, which broadens the results by including papers that match any of the keywords.
3.3) “AND” + “OR” logic
Now, a combination of AND and OR will be conducted as follows.
Next, I searched for the keywords Soybean OR Yield AND maize OR nitrogen in the title, and Grain OR Pod AND Kernel OR Weight in the abstract, restricting the publication years to 2020–2023. I ran the following code.
reference= scopusmining(
title_blocks= list(
list(terms= c("Soybean", "Yield"), logic= "OR"),
list(terms= c("Maize", "Nitrogen"), logic= "OR")),
abskey_blocks= list(
list(terms= c("Grain", "Pod"), logic= "OR"),
list(terms= c("Kernel", "Weight"), logic= "OR")),
years= 2020:2023,
dedupe= FALSE
)
The search query used in this search is presented as follows.
TITLE(("Soybean" OR "Yield") AND ("Maize" OR "Nitrogen")) AND TITLE-ABS-KEY(("Grain" OR "Pod") AND ("Kernel" OR "Weight")) AND DOCTYPE(AR) AND SRCTYPE(j)
3.4) Repeated searches will be ignored
If keywords are duplicated, it can be ignored using the following code.
reference= scopusmining(
title_blocks= list(
list(terms= c("Soybean", "Yield", "Biomass"), logic= "OR")),
abskey_blocks= list(
list(terms= c("Soybean", "Yield", "Pod"), logic= "OR")),
years= 2020:2023,
dedupe= TRUE
)
The search query used in this search is presented as follows.
TITLE(("Soybean" OR "Yield" OR "Biomass")) AND TITLE-ABS-KEY("Pod") AND DOCTYPE(AR) AND SRCTYPE(j)

We aim to develop open-source code for agronomy ([email protected])
© 2022 – 2025 https://agronomy4future.com – All Rights Reserved.
Last Updated: 12/22/2025