In this session, I will introduce the method of calculating the Best Linear Unbiased Estimator (BLUE). Instead of simply listing formulas as many websites do to explain BLUE, this post aims to help readers understand the process of calculating BLUE with an actual dataset using R.
I have the following data.
df= structure(list(location = c("Cordoba", "Cordoba", "Cordoba", "Cordoba", "Cordoba", "Cordoba", "Cordoba", "Cordoba", "Cordoba", "Cordoba", "Cordoba", "Cordoba", "Granada", "Granada", "Granada", "Granada", "Granada", "Granada", "Granada", "Granada", "Granada", "Granada", "Granada", "Granada", "Granada", "Granada", "Granada", "Granada", "Leon", "Leon", "Leon", "Leon", "Leon", "Leon", "Leon", "Leon", "Leon", "Leon", "Leon", "Leon"), trt = c(0L, 24L, 36L, 48L, 0L, 24L, 36L, 48L, 0L, 24L, 36L, 48L, 0L, 24L, 36L, 48L, 0L, 24L, 36L, 48L, 0L, 24L, 36L, 48L, 0L, 24L, 36L, 48L, 0L, 24L, 36L, 48L, 0L, 24L, 36L, 48L, 0L, 24L, 36L, 48L), block = structure(c(2L, 2L, 2L, 2L, 3L, 3L, 3L, 3L, 4L, 4L, 4L, 4L, 1L, 1L, 1L, 1L, 2L, 2L, 2L, 2L, 3L, 3L, 3L, 3L, 4L, 4L, 4L, 4L, 1L, 1L, 1L, 1L, 3L, 3L, 3L, 3L, 4L, 4L, 4L, 4L), levels = c("1", "2", "3", "4"), class = "factor"), yield = c(780, 1280, 1520, 1130, 790, 1300, 1525, 1140, 800, 10, 15, 11, 10, 1490, 1790, 1345, 1066, 1510, 1700, 1355, 948, 1560, 1830, 1368, 960, 1550, 1866, 1356, 800, 1250, 1550, 910, 663.6, 1260, 1480, 900, 750, 1195, 1525, 900)), row.names = c(NA, -40L), class = "data.frame")
The dataset comprises three different locations involving four sulfur experiments organized into four blocks for crop yield; however, some blocks were missing in Cordoba and Leon.
df$block= as.factor(df$block) print(df) location trt block yield 1 Cordoba 0 2 780.0 2 Cordoba 24 2 1280.0 3 Cordoba 36 2 1520.0 4 Cordoba 48 2 1130.0 5 Cordoba 0 3 790.0 6 Cordoba 24 3 1300.0 7 Cordoba 36 3 1525.0 8 Cordoba 48 3 1140.0 9 Cordoba 0 4 800.0 10 Cordoba 24 4 10.0 11 Cordoba 36 4 15.0 12 Cordoba 48 4 11.0 13 Granada 0 1 10.0 14 Granada 24 1 1490.0 15 Granada 36 1 1790.0 16 Granada 48 1 1345.0 17 Granada 0 2 1066.0 18 Granada 24 2 1510.0 19 Granada 36 2 1700.0 20 Granada 48 2 1355.0 21 Granada 0 3 948.0 22 Granada 24 3 1560.0 23 Granada 36 3 1830.0 24 Granada 48 3 1368.0 25 Granada 0 4 960.0 26 Granada 24 4 1550.0 27 Granada 36 4 1866.0 28 Granada 48 4 1356.0 29 Leon 0 1 800.0 30 Leon 24 1 1250.0 31 Leon 36 1 1550.0 32 Leon 48 1 910.0 33 Leon 0 3 663.6 34 Leon 24 3 1260.0 35 Leon 36 3 1480.0 36 Leon 48 3 900.0 37 Leon 0 4 750.0 38 Leon 24 4 1195.0 39 Leon 36 4 1525.0 40 Leon 48 4 900.0
Block 1 is missing from the Cordoba data, and Block 2 is missing from Leon. Do you think it is acceptable to use this unbalanced data as is, or should we apply a method to compensate for the imbalance?
To help you understand, let’s use a very simple analogy.
Three Students taking a 4-Set Exam. Imagine three students are taking an exam. The exam consists of four different sets.
Student A: Completed both the easy Set 1 and the difficult Set 2 (Average Score: 50)
Student B: Completed only the difficult Set 2 (Average Score: 30)
If we only look at the simple average, Student A (50) seems smarter than Student B (30). We might wrongly conclude that Student B is a poor student.
The BLUE (Mixed Model) Approach
The Mixed Model doesn't just look at the final score; it looks at the context (i.e., Random effect).
The model analyzes all the data and realizes, "Wait, everyone scored 20 points lower on Set 2 because it was much harder." Since Student B only tackled the most difficult set, the model adjusts the score: "If Student B had also taken the easy set, they likely would have scored a 50 as well." The model concludes that the actual ability (Fixed Effect) of Student A and Student B is actually similar.
# Fixed Effect: The actual underlying factor we want to measure (e.g., student ability or crop yield).
# Random/Block Effect: The environmental noise or context that can bias the results (e.g., difficulty of the exam set or soil quality of a specific block).
# Adjusted Means: The "fair" scores calculated after removing the bias of the blocks.
Consider a dataset investigating crop yields in three regions of Spain under sulfur fertilizer treatments. The experiment was conducted using four repetitions as blocks. In this model, location is defined as a fixed factor, while block are defined as random factors (For now, let’s disregard sulfur, assuming its effect is consistent across all locations.).
In such a mixed-effects model, the Best Linear Unbiased Estimator (BLUE) is used to estimate the fixed effects, while the Best Linear Unbiased Prediction (BLUP) is used to predict the random effects. I will now demonstrate how to calculate both the BLUEs and BLUPs using R.

To calculate BLUE using R
I will configure the model. Location is set as a fixed factor, while block are set as a random factor. The model configuration is as follows.
if(!require(lme4)) install.packages("lme4")
if(!require(lmerTest)) install.packages("lmerTest")
library(lme4)
library(lmerTest)
model=lmer(yield ~ 0 + location + (1|block), data=df)
The reason I set up the intercept as 0 is because I want the model to directly estimate the mean yield (BLUE) for each specific location, rather than calculating the differences relative to a reference region.
Why suppressing the Intercept?
When configuring the model asyield ~ 0 + location + (1|block), we are using what is known as a "Cell-Means Model." This setup is chosen for the following reasons:
1. Direct Estimation of Absolute Means (BLUEs)
In a default model (yield ~ location), R treats the first alphabetical location as a reference level (intercept) and shows other locations as differences from that baseline. By adding 0 (or -1), we remove this baseline. This forces the model to calculate the absolute mean yield for every location directly. These resulting coefficients are the BLUEs (Best Linear Unbiased Estimators) for each specific region.
2. Efficiency in Reporting and Interpretation
If the primary goal of your study is to report the actual productivity of each of the three Spanish regions, the 0 + location approach is much more efficient. You can extract the results using fixef(model) and report them immediately without having to manually add the intercept to the slope coefficients.
3. Objective Comparison Without Bias
Treating one specific region as a "standard" (intercept) can sometimes lead to biased interpretations in comparative studies. Suppressing the intercept ensures that each region is treated as an independent fixed factor, providing a more balanced and objective view of the geographic yield distribution.
4. Handling Unbalanced Data through BLUE
As we discussed, when data is unbalanced (e.g., some blocks are missing in certain regions), the 0 + location setup ensures that the estimate for each region is adjusted for the random effects of the blocks. The model looks at the block performance across other regions to "correct" the missing or skewed data in a specific region, providing the most "unbiased" version of that region's mean.
In summary: You use 0 + location because you want the model outputs to represent the true, adjusted average yield (BLUE) of each location, rather than just showing how much they differ from a reference point.
Let’s see the output of the model.
summary(model)
Linear mixed model fit by REML. t-tests use Satterthwaite's method [
lmerModLmerTest]
Formula: yield ~ 0 + location + (1 | block)
Data: df
REML criterion at convergence: 565.8
Scaled residuals:
Min 1Q Median 3Q Max
-2.9351 -0.6253 0.1773 0.6523 1.3901
Random effects:
Groups Name Variance Std.Dev.
block (Intercept) 15481 124.4
Residual 199021 446.1
Number of obs: 40, groups: block, 4
Fixed effects:
Estimate Std. Error df t value Pr(>|t|)
locationCordoba 846.05 145.85 11.73 5.801 9.27e-05 ***
locationGranada 1356.50 127.71 9.25 10.622 1.73e-06 ***
locationLeon 1123.10 145.85 11.73 7.701 6.38e-06 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Correlation of Fixed Effects:
lctnCr lctnGr
locatinGrnd 0.208
locationLen 0.167 0.208
In the lmer output, the three values listed under the Estimate column in the Fixed effects section (846.05, 1356.50, 1123.10) are indeed the BLUE (Best Linear Unbiased Estimator) values for each location.
Why are these values called "BLUE"?
□ Best and Unbiased: These are not just simple arithmetic means of the collected data. They are the most "unbiased" optimal estimates, obtained by statistically adjusting (removing) the noise from random effects (Block) included in the model.
□ Location-Specific: Because you used the 0 + location syntax, these values represent the pure adjusted mean for each specific region, rather than the difference relative to a reference baseline.
# Fixed Effects: Population-level average regression coefficients (Named Vector)
print(data.frame(fixef(model)))
fixef.model.
locationCordoba 846.0555
locationGranada 1356.5000
locationLeon 1123.0997
Let’s see the arithmetic mean of each location.
if(!require(dplyr)) install.packages("dplyr")
library(dplyr)
structure = df %>%
group_by(location) %>%
dplyr::summarize(
across(
.cols= yield,
.fns= list(
Mean= ~mean(., na.rm= TRUE),
n= ~length(.),
se= ~sd(., na.rm= TRUE) / sqrt(length(.)))),
.groups= "drop") %>%
as.data.frame()
print(structure)
location yield_Mean yield_n yield_se
1 Cordoba 858.4167 12 164.48544
2 Granada 1356.5000 16 114.23204
3 Leon 1098.6333 12 91.43936
You can see the arithmetic (simple) mean is different from BLUE.
| Location | Simple mean | BLUE |
| Cordoba | 858.4167 | 846.05 |
| Granada | 1356.5000 | 1356.50 |
| Leon | 1098.6333 | 1123.10 |
While the simple mean only averages the available observed data, BLUE provides a more balanced estimate by accounting for both missing blocks and localized yield anomalies. For Cordoba, the model identifies that the exceptionally low yields in Block 4 are a ‘block-specific’ issue rather than a regional one, and it also leverages data from other regions to compensate for the poor/missing information in Block 1.
In the case of Leon, which lacks any data for Block 2, the model uses the performance of Block 2 in Granada and Cordoba to estimate what Leon’s yield would have been. Since Block 2 generally shows higher yields, BLUE adjusts Leon’s estimate upward to reflect its true potential. By ‘filling in’ these gaps and adjusting for the noise in Cordoba’s Block 4, BLUE offers a far more accurate representation of each region’s productivity than the biased simple mean.
For Granada, where the experimental design was perfectly balanced across all blocks and treatments, the simple mean and the BLUE are identical. This confirms that there was no regional bias or missing information to be adjusted in Granada, unlike Cordoba and Leon, where BLUE corrected for missing blocks and localized yield anomalies

Now let’s calculate the prediction values from the model and add these prediction values into the existing data.
df$prediction= predict (model, re.form = NA)
Now let’s calculate the average of these prediction values.
if(!require(dplyr)) install.packages("dplyr")
library(dplyr)
summary = df %>%
group_by(location) %>%
dplyr::summarize(
across(
.cols= prediction,
.fns= list(
Mean= ~mean(., na.rm= TRUE),
n= ~length(.),
se= ~sd(., na.rm= TRUE) / sqrt(length(.)))),
.groups= "drop") %>%
as.data.frame()
print(summary) location prediction_Mean prediction_n prediction_se 1 Cordoba 846.0555 12 0 2 Granada 1356.5000 16 0 3 Leon 1123.0997 12 0
Did you find that the mean of prediction values from the model is the same as BLUE and also standard error becomes 0?
print(df) location trt block yield prediction 1 Cordoba 0 2 780.0 846.0555 2 Cordoba 24 2 1280.0 846.0555 3 Cordoba 36 2 1520.0 846.0555 4 Cordoba 48 2 1130.0 846.0555 5 Cordoba 0 3 790.0 846.0555 6 Cordoba 24 3 1300.0 846.0555 7 Cordoba 36 3 1525.0 846.0555 8 Cordoba 48 3 1140.0 846.0555 9 Cordoba 0 4 800.0 846.0555 10 Cordoba 24 4 10.0 846.0555 11 Cordoba 36 4 15.0 846.0555 12 Cordoba 48 4 11.0 846.0555 13 Granada 0 1 10.0 1356.5000 14 Granada 24 1 1490.0 1356.5000 15 Granada 36 1 1790.0 1356.5000 16 Granada 48 1 1345.0 1356.5000 17 Granada 0 2 1066.0 1356.5000 18 Granada 24 2 1510.0 1356.5000 19 Granada 36 2 1700.0 1356.5000 20 Granada 48 2 1355.0 1356.5000 21 Granada 0 3 948.0 1356.5000 22 Granada 24 3 1560.0 1356.5000 23 Granada 36 3 1830.0 1356.5000 24 Granada 48 3 1368.0 1356.5000 25 Granada 0 4 960.0 1356.5000 26 Granada 24 4 1550.0 1356.5000 27 Granada 36 4 1866.0 1356.5000 28 Granada 48 4 1356.0 1356.5000 29 Leon 0 1 800.0 1123.0997 30 Leon 24 1 1250.0 1123.0997 31 Leon 36 1 1550.0 1123.0997 32 Leon 48 1 910.0 1123.0997 33 Leon 0 3 663.6 1123.0997 34 Leon 24 3 1260.0 1123.0997 35 Leon 36 3 1480.0 1123.0997 36 Leon 48 3 900.0 1123.0997 37 Leon 0 4 750.0 1123.0997 38 Leon 24 4 1195.0 1123.0997 39 Leon 36 4 1525.0 1123.0997 40 Leon 48 4 900.0 1123.0997
Using predict(model, re.form = NA) means ŷ = Fixed Effect (Location). It ignores which block the data came from. It asks: “What is the general potential of this location if we strip away the luck of having a good or bad block? All values at each location will get the exact same value. The standard error of these values is 0 because you are looking at a single constant estimate (the BLUE).
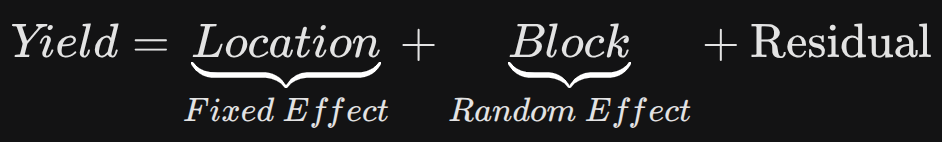
On the other hand, using predict(model) means ŷ = Fixed Effect (Location) + Random Effect (Block). It accounts for the specific conditions of that data point. It asks: “Based on what we know about this specific block and this specific treatment, what was the most likely yield?” If you include random factors using df$prediction2 = predict(model), it means you are calculating the ‘Subject-level’ (or group-level) prediction. This value represents the expected yield for a specific observation by combining the regional average (BLUE) with the specific deviations of the block (BLUPs) associated with that row.
# Random Effects: Group-specific deviations/offsets from the population average (List of Dataframes) print(data.frame(ranef(model))) grpvar term grp condval condsd 1 block (Intercept) 1 -37.08357 97.68736 2 block (Intercept) 2 73.39903 97.68736 3 block (Intercept) 3 58.81900 89.48238 4 block (Intercept) 4 -95.13446 89.48238
Look at the prediction2 values in the data below. prediction2 represents ŷ = Fixed Effect (Location) + Random Effect (Block). If I subtract the condval (Conditional Value) of each block from prediction2, the resulting value is the population-level prediction (Fixed Effect only). For example, 919.4545 – 73.39903 ≈ 846.055.
df$prediction2=predict (model) print(df) location trt block yield prediction prediction2 1 Cordoba 0 2 780.0 846.0555 919.4545 2 Cordoba 24 2 1280.0 846.0555 919.4545 3 Cordoba 36 2 1520.0 846.0555 919.4545 4 Cordoba 48 2 1130.0 846.0555 919.4545 5 Cordoba 0 3 790.0 846.0555 904.8745 6 Cordoba 24 3 1300.0 846.0555 904.8745 7 Cordoba 36 3 1525.0 846.0555 904.8745 8 Cordoba 48 3 1140.0 846.0555 904.8745 9 Cordoba 0 4 800.0 846.0555 750.9210 10 Cordoba 24 4 10.0 846.0555 750.9210 11 Cordoba 36 4 15.0 846.0555 750.9210 12 Cordoba 48 4 11.0 846.0555 750.9210 13 Granada 0 1 10.0 1356.5000 1319.4164 14 Granada 24 1 1490.0 1356.5000 1319.4164 15 Granada 36 1 1790.0 1356.5000 1319.4164 16 Granada 48 1 1345.0 1356.5000 1319.4164 17 Granada 0 2 1066.0 1356.5000 1429.8990 18 Granada 24 2 1510.0 1356.5000 1429.8990 19 Granada 36 2 1700.0 1356.5000 1429.8990 20 Granada 48 2 1355.0 1356.5000 1429.8990 21 Granada 0 3 948.0 1356.5000 1415.3190 22 Granada 24 3 1560.0 1356.5000 1415.3190 23 Granada 36 3 1830.0 1356.5000 1415.3190 24 Granada 48 3 1368.0 1356.5000 1415.3190 25 Granada 0 4 960.0 1356.5000 1261.3655 26 Granada 24 4 1550.0 1356.5000 1261.3655 27 Granada 36 4 1866.0 1356.5000 1261.3655 28 Granada 48 4 1356.0 1356.5000 1261.3655 29 Leon 0 1 800.0 1123.0997 1086.0161 30 Leon 24 1 1250.0 1123.0997 1086.0161 31 Leon 36 1 1550.0 1123.0997 1086.0161 32 Leon 48 1 910.0 1123.0997 1086.0161 33 Leon 0 3 663.6 1123.0997 1181.9187 34 Leon 24 3 1260.0 1123.0997 1181.9187 35 Leon 36 3 1480.0 1123.0997 1181.9187 36 Leon 48 3 900.0 1123.0997 1181.9187 37 Leon 0 4 750.0 1123.0997 1027.9652 38 Leon 24 4 1195.0 1123.0997 1027.9652 39 Leon 36 4 1525.0 1123.0997 1027.9652 40 Leon 48 4 900.0 1123.0997 1027.9652
In the Cordoba region, Block 4 exhibited exceptionally low observed yields (10.0–15.0). However, the linear mixed-effects model adjusted these values; when calculating conditional predictions; predict(model), the estimate for Block 4 rose to 750.92. This adjustment occurs because the model accounts for the overall block-level variance through Best Linear Unbiased Predictors (BLUPs). Notably, the mean of these conditional predictions remains mathematically identical to the simple arithmetic mean of the observed data because the BLUPs are centered around the group mean to minimize residuals.
summary = df %>%
group_by(location) %>%
dplyr::summarize(
across(
.cols= c(prediction, prediction2),
.fns= list(
Mean= ~mean(., na.rm= TRUE),
n= ~length(.),
se= ~sd(., na.rm= TRUE) / sqrt(length(.)))),
.groups= "drop") %>%
as.data.frame()
summary= subset(summary,
select=c(location,prediction_Mean,prediction_se,
prediction2_Mean,prediction2_se))
print(summary)
location prediction_Mean prediction_se prediction2_Mean prediction2_se
1 Cordoba 846.0555 0 858.4167 22.98831
2 Granada 1356.5000 0 1356.5000 17.92245
3 Leon 1123.0997 0 1098.6333 19.14033
To report the Best Linear Unbiased Estimators (BLUEs)—the true adjusted means—one must instead utilize the fixed-effects estimates by excluding random factors adding re.form = NA. The standard error for these BLUEs represents the uncertainty of the fixed-effect estimate and should be obtained from the model’s fixed-effects covariance matrix or via specialized packages like emmeans, rather than calculating the standard deviation of the predicted constants.

How to calculate standard error of BLUE?
The standard error (SE) of the Best Linear Unbiased Estimators (BLUEs) was calculated using the emmeans() package for the following reasons
First, unlike simple arithmetic means, emmeans()
Second, while the standard deviation of raw predictions often fails to reflect model uncertainty, emmeans() derives the SE directly from the fixed-effects covariance matrix.
Finally, the package utilizes the Kenward-Roger degrees-of-freedom approximation, which is the gold standard for providing robust SEs and confidence intervals in linear mixed-effects models, ensuring that the precision of each regional estimate is accurately represented regardless of sample size differences.
if(!require(emmeans)) install.packages("emmeans")
library(emmeans)
model= lmer(yield ~ 0 + location + (1|block), data = df)
output= emmeans(model, "location")
print(output) location emmean SE df lower.CL upper.CL Cordoba 846 151 11.30 515 1177 Granada 1356 128 8.88 1067 1646 Leon 1123 151 11.30 792 1455 Degrees-of-freedom method: kenward-roger Confidence level used: 0.95

We aim to develop open-source code for agronomy ([email protected])
© 2022 – 2025 https://agronomy4future.com – All Rights Reserved.
Last Updated: 04/07/2026