In my previous posts, I introduced how to divide panels in one figure using facet_wrap(). Today, I’ll introduce how to add separate text to panels.
First, let’s make sure we have the required packages installed.
if(!require(ggplot2)) install.packages("ggplot2")
if(!require(dplyr)) install.packages("dplyr")
if(!require(tidyr)) install.packages("tidyr")
library (ggplot2)
library (dplyr)
library (tidyr)
I’ll create a dataset as shown below:
Nitrogen= c(rep("N0", 5), rep("N1", 5))
Cultivar_1= c(50,49,48,47,46,60,62,63,64,62)
Cultivar_2= c(55,57,56,55,54,65,66,67,64,63)
Cultivar_3= c(60,62,63,65,59,60,59,57,56,58)
dataA= data.frame(Nitrogen, Cultivar_1, Cultivar_2, Cultivar_3)
dataA
Nitrogen Cultivar_1 Cultivar_2 Cultivar_3
1 N0 50 55 60
2 N0 49 57 62
3 N0 48 56 63
4 N0 47 55 65
5 N0 46 54 59
6 N1 60 65 60
7 N1 62 66 59
8 N1 63 67 57
9 N1 64 64 56
10 N1 62 63 58
Next, I’ll reshape the dataset into columns to facilitate data analysis.
df= data.frame(
dataA %>%
pivot_longer(
cols=c(Cultivar_1, Cultivar_2, Cultivar_3),
names_to="Genotype", values_to="Yield")
)
df
Nitrogen Genotype Yield
N0 Cultivar_1 50
N0 Cultivar_2 55
N0 Cultivar_3 60
N0 Cultivar_1 49
N0 Cultivar_2 57
N0 Cultivar_3 62
.
.
.
And then, I’ll summarize this data using descriptive statistics.
dataB= data.frame(df %>%
group_by(Genotype, Nitrogen) %>%
dplyr::summarize(across(c(Yield),
.fns = list(Mean = mean,
SD = sd,
n = length,
se = ~ sd(.)/sqrt(length(.))))))
dataB
Genotype Nitrogen Yield_Mean Yield_SD Yield_n Yield_se
Cultivar_1 N0 48.0 1.581139 5 0.7071068
Cultivar_1 N1 62.2 1.483240 5 0.6633250
Cultivar_2 N0 55.4 1.140175 5 0.5099020
Cultivar_2 N1 65.0 1.581139 5 0.7071068
Cultivar_3 N0 61.8 2.387467 5 1.0677078
Cultivar_3 N1 58.0 1.581139 5 0.7071068
Finally, I’ll create a graph to visualize the summarized data.
ggplot (data=dataB, aes(x=Genotype, y=Yield_Mean, fill=Nitrogen)) +
geom_bar(stat="identity",position="dodge") +
geom_errorbar(aes(ymin= Yield_Mean-Yield_se, ymax= Yield_Mean+Yield_se),
position=position_dodge(0.9), width=0.2) +
scale_fill_manual(values= c("Gray","Dark green")) +
scale_y_continuous(breaks= seq(0, 100, 20), limits= c(0, 100))+
facet_wrap(~Nitrogen) +
labs(x="Cultivar", y="Yield") +
theme_classic(base_size=18, base_family="serif")+
theme(legend.position='none',
legend.title=element_blank(),
legend.key=element_rect(color="white", fill="white"),
legend.text=element_text(family="serif", face="plain",
size=15, color= "Black"),
legend.background=element_rect(fill="white"),
axis.line=element_line(linewidth=0.5, colour="black"),
strip.background=element_rect(color="white",
linewidth=0.5,linetype="solid"))
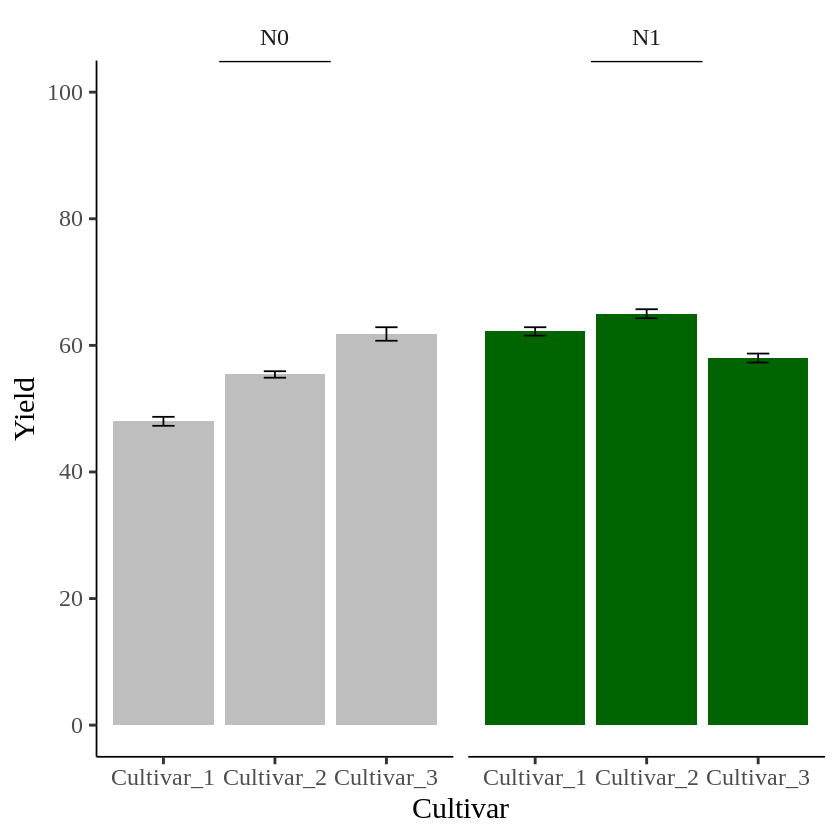
In this graph, I want to indicate statistical significance with asterisks (***) in each panel. First, I’ll add asterisks in the N0 panel.
ggplot (data=dataB, aes(x=Genotype, y=Yield_Mean, fill=Nitrogen)) +
geom_bar(stat="identity",position="dodge") +
geom_errorbar(aes(ymin= Yield_Mean-Yield_se, ymax= Yield_Mean+Yield_se),
position=position_dodge(0.9), width=0.2) +
scale_fill_manual(values= c("Gray","Dark green")) +
###
geom_text(aes(family="serif",fontface=6), x=1.5, y=80, label="***", size=7, col="red") +
###
scale_y_continuous(breaks= seq(0, 100, 20), limits= c(0, 100))+
facet_wrap(~Nitrogen) +
annotate("segment", x=1.5, xend=2.5, y=Inf,yend=Inf, color="black", lwd=1)+
labs(x="Cultivar", y="Yield") +
theme_classic(base_size=18, base_family="serif")+
theme(legend.position='none',
legend.title=element_blank(),
legend.key=element_rect(color="white", fill="white"),
legend.text=element_text(family="serif", face="plain",
size=15, color= "Black"),
legend.background=element_rect(fill="white"),
axis.line=element_line(linewidth=0.5, colour="black"),
strip.background=element_rect(color="white",
linewidth=0.5,linetype="solid"))
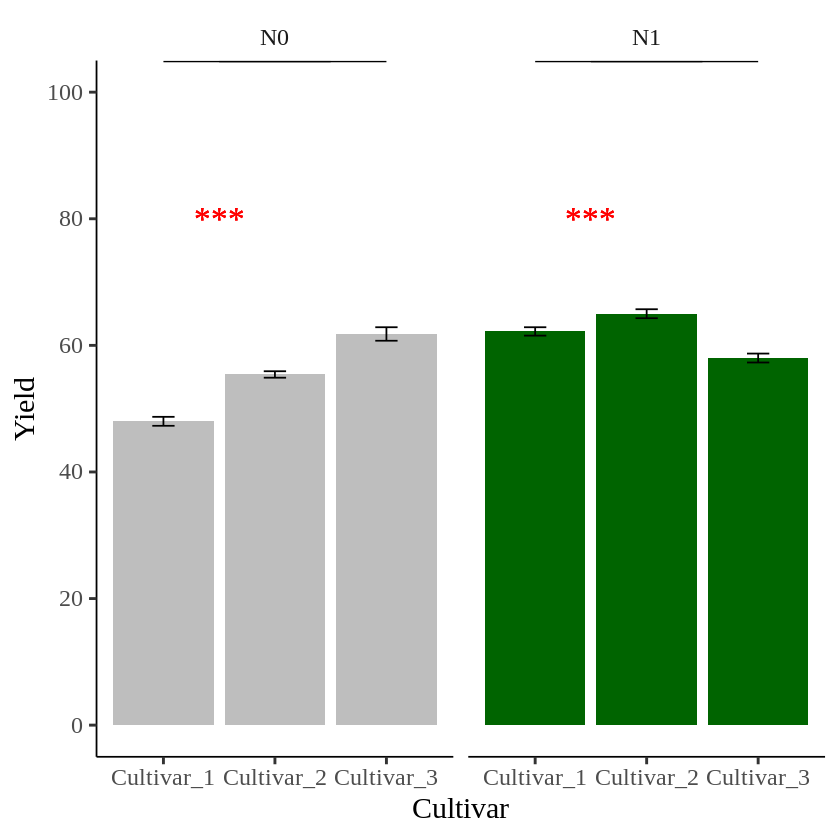
The problem is that *** is added to both panels. I want a different symbol for each panel.

to create data table to insert text to specific panel
I’ll create a data table.
text1= data.frame(Nitrogen="N0", STAT= "***", Genotype="Cultivar_2", Yield=70) text1 Nitrogen STAT Genotype Yield N0 *** Cultivar_2 70
I’ll add this data table to the geom_text().
geom_text(data=text1, aes(family="serif", x=Genotype, y=Yield, label=STAT), size=5, color="red")
Let’s create a graph again, this time to visualize the data.
ggplot (data=dataB, aes(x=Genotype, y=Yield_Mean, fill=Nitrogen)) +
geom_bar(stat="identity",position="dodge") +
geom_errorbar(aes(ymin= Yield_Mean-Yield_se, ymax= Yield_Mean+Yield_se),
position=position_dodge(0.9), width=0.2) +
scale_fill_manual(values= c("Gray","Dark green")) +
###
geom_text(data=text1, aes(family="serif", x=Genotype, y=Yield, label=STAT), size=5, color="red") +
###
scale_y_continuous(breaks= seq(0, 100, 20), limits= c(0, 100))+
facet_wrap(~Nitrogen) +
annotate("segment", x=1.5, xend=2.5, y=Inf,yend=Inf, color="black", lwd=1)+
labs(x="Cultivar", y="Yield") +
theme_classic(base_size=18, base_family="serif")+
theme(legend.position='none',
legend.title=element_blank(),
legend.key=element_rect(color="white", fill="white"),
legend.text=element_text(family="serif", face="plain",
size=15, color= "Black"),
legend.background=element_rect(fill="white"),
axis.line=element_line(linewidth=0.5, colour="black"),
strip.background=element_rect(color="white",
linewidth=0.5,linetype="solid"))
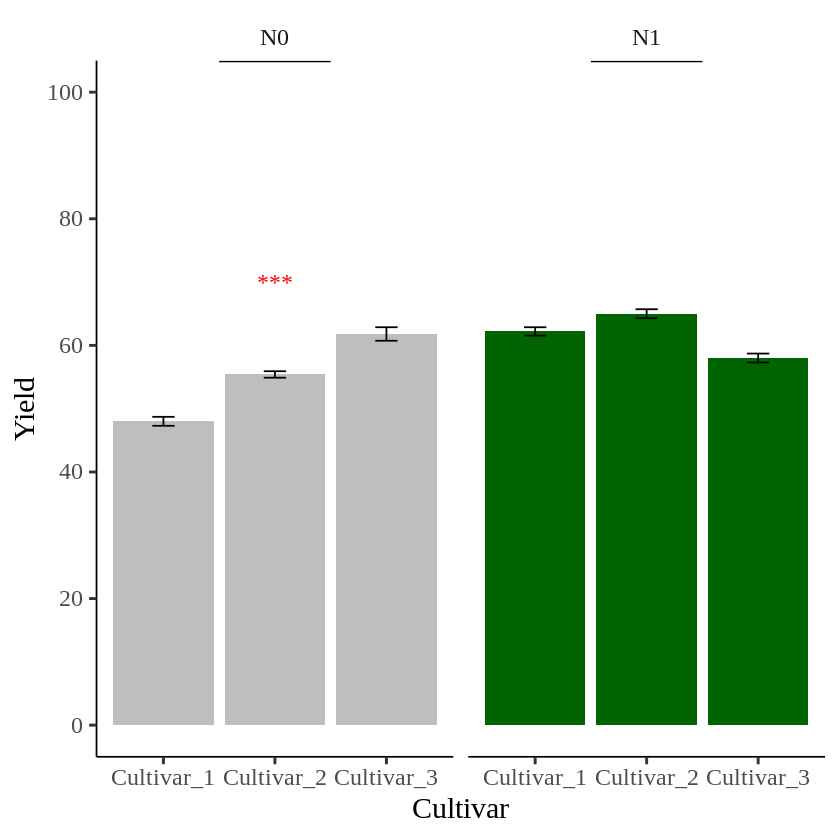
I’ll create another data table to add text in the second panel.
text2= data.frame(Nitrogen="N1", STAT= "**", Genotype="Cultivar_1", Yield=80)
geom_text(data=text2, aes(family="serif", x=Genotype, y=Yield, label=STAT), size=5, color="red")
##
ggplot (data=dataB, aes(x=Genotype, y=Yield_Mean, fill=Nitrogen)) +
geom_bar(stat="identity",position="dodge") +
geom_errorbar(aes(ymin= Yield_Mean-Yield_se, ymax= Yield_Mean+Yield_se),
position=position_dodge(0.9), width=0.2) +
scale_fill_manual(values= c("Gray","Dark green")) +
###
geom_text(data=text1, aes(family="serif", x=Genotype, y=Yield, label=STAT), size=5, color="red") +
geom_text(data=text2, aes(family="serif", x=Genotype, y=Yield, label=STAT), size=5, color="red") +
###
scale_y_continuous(breaks= seq(0, 100, 20), limits= c(0, 100))+
facet_wrap(~Nitrogen) +
annotate("segment", x=1.5, xend=2.5, y=Inf,yend=Inf, color="black", lwd=1)+
labs(x="Cultivar", y="Yield") +
theme_classic(base_size=18, base_family="serif")+
theme(legend.position='none',
legend.title=element_blank(),
legend.key=element_rect(color="white", fill="white"),
legend.text=element_text(family="serif", face="plain",
size=15, color= "Black"),
legend.background=element_rect(fill="white"),
axis.line=element_line(linewidth=0.5, colour="black"),
strip.background=element_rect(color="white",
linewidth=0.5,linetype="solid"))
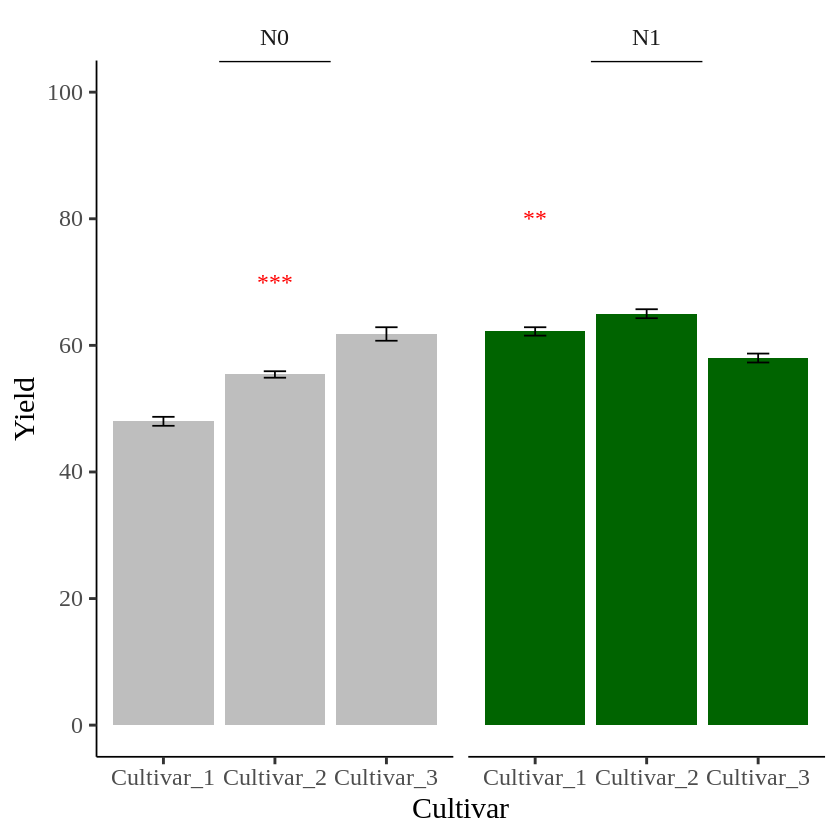
Now, different asterisks (***) are indicated in each panel.
full code: https://github.com/agronomy4future/r_code/blob/main/How_to_add_separate_text_to_panels_divided_by_facet_wrap()_in_R.ipynb

We aim to develop open-source code for agronomy ([email protected])
© 2022 – 2025 https://agronomy4future.com – All Rights Reserved.
Last Updated: 02/27/2024