if(!require(remotes)) install.packages("remotes")
library(remotes)
if (!requireNamespace("rnmodel", quietly = TRUE)) {
remotes::install_github("agronomy4future/rnmodel", force= TRUE)
}
library(rnmodel)
if(!require(readr)) install.packages("readr")
library(readr)
github= paste0("https://raw.githubusercontent.com/agronomy4future/",
"raw_data_practice/refs/heads/main/",
"nitrogen_level_crop_yield.csv")
df= data.frame(read_csv(url(github),show_col_types = FALSE))
output= rnmodel(data= df, genotype_by= c("Genotype","Replicate"),
environment="Nitrogen_Level",
response="Yield",
env_numeric = NULL)
result=output$reaction_norm
set.seed(100)
print(result[sample(nrow(result),5),])
Reaction Norm Model Output (Biological Terms):
Genotype Replicate baseline plasticity baseline_se plasticity_se
503 CV2 3 3.22400 0.0063072 0.3199945 0.002612744
2035 CV5 35 2.95600 0.0193400 0.3123171 0.002550059
470 CV1 470 3.31600 -0.0008200 0.1552482 0.001267596
1990 CV4 490 3.96964 0.0041024 0.3560298 0.002906971
1540 CV4 40 3.50714 0.0057524 0.4093860 0.003342623
Fig1=ggplot(data=result, aes(x=Genotype, y=plasticity, fill=Genotype)) +
geom_boxplot() +
scale_color_manual(values=c("cadetblue","darkorchid4","orange3",
"skyblue4","wheat4"))+
scale_fill_manual(values=c("cadetblue","darkorchid4","orange3",
"skyblue4","wheat4"))+
labs(x="Genotype", y="Plascitiy (slope)") +
theme_classic(base_size=15, base_family="serif") +
theme(legend.position="none",
legend.title=element_blank(),
legend.key=element_rect(color="white", fill=alpha(0.5)),
legend.text=element_text(family="serif", face="plain",
size=13, color="black"),
legend.background= element_rect(fill=alpha(0.5)),
axis.title.y = element_text(margin= margin(r= 15)),
panel.border= element_rect(color="black", fill=NA, linewidth=0.5),
axis.line= element_line(linewidth=0.5, colour="black"),
strip.background=element_rect(color="white",
linewidth=0.5, linetype="solid"))
options(repr.plot.width=5.5, repr.plot.height=5)
print(Fig1)
ggsave("Fig1.png", plot= Fig1, width=5.5, height= 5, dpi= 300)
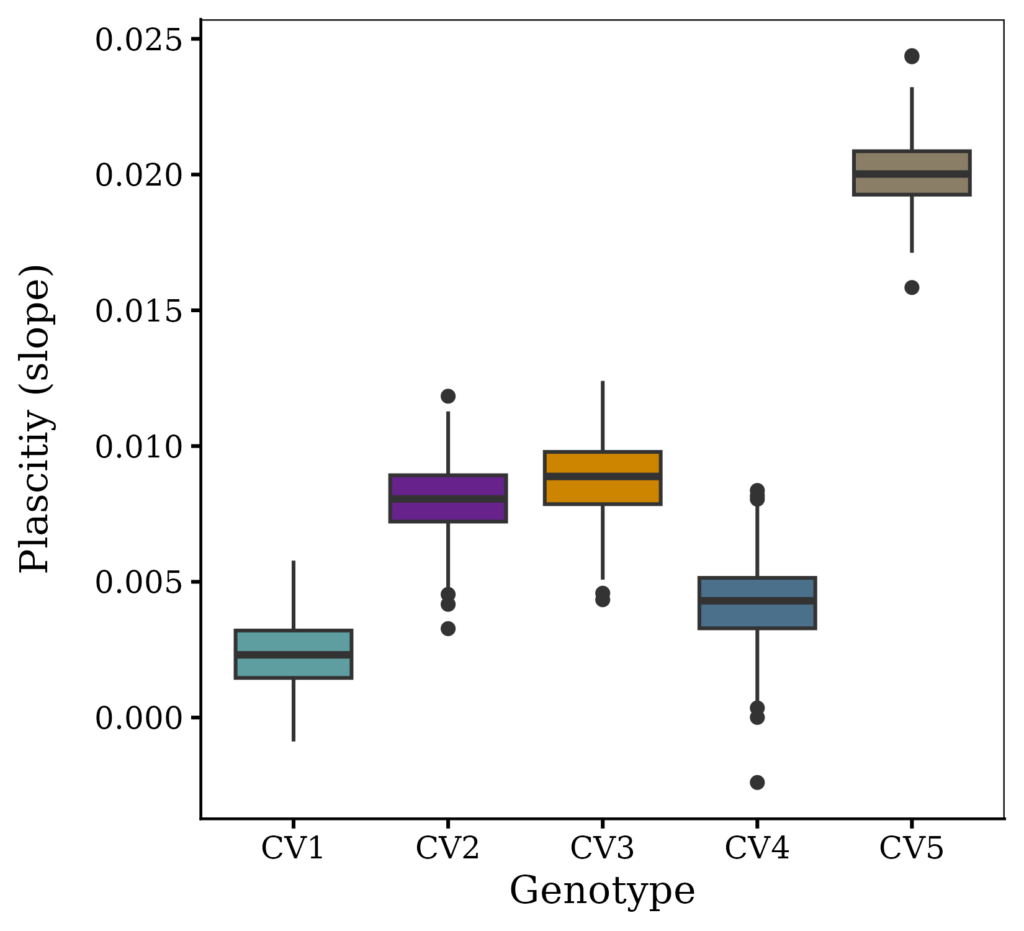
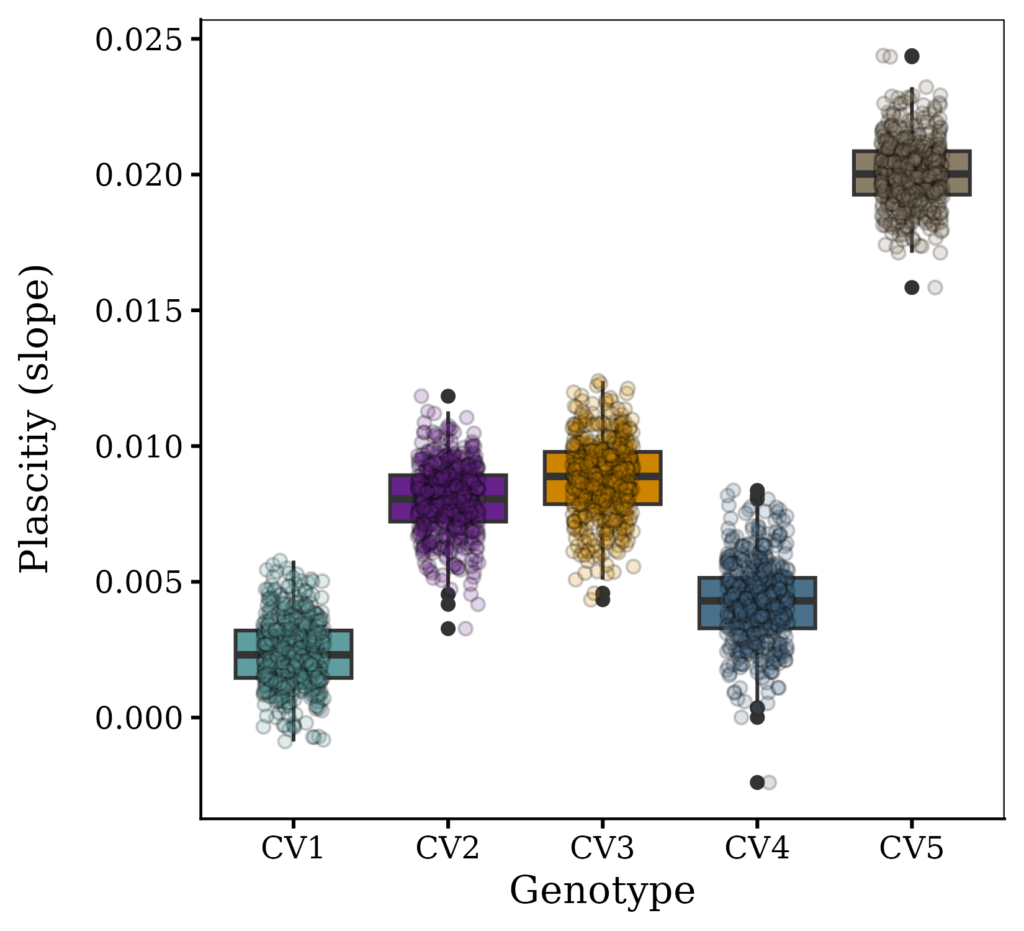
Fig1=ggplot(data=result, aes(x=Genotype, y=plasticity, fill=Genotype)) +
geom_boxplot() +
geom_jitter(data=result, aes(x= Genotype, y= plasticity,
fill=Genotype, shape=Genotype), width=0.2, alpha=0.2, size=2) +
scale_color_manual(values=c("cadetblue","darkorchid4","orange3",
"skyblue4","wheat4"))+
scale_fill_manual(values=c("cadetblue","darkorchid4","orange3",
"skyblue4","wheat4"))+
scale_shape_manual(values=c(21,21,21,21,21))+
labs(x="Genotype", y="Plascitiy (slope)") +
theme_classic(base_size=15, base_family="serif") +
theme(legend.position="none",
legend.title=element_blank(),
legend.key=element_rect(color="white", fill=alpha(0.5)),
legend.text=element_text(family="serif", face="plain",
size=13, color="black"),
legend.background= element_rect(fill=alpha(0.5)),
axis.title.y = element_text(margin= margin(r= 15)),
panel.border= element_rect(color="black", fill=NA, linewidth=0.5),
axis.line= element_line(linewidth=0.5, colour="black"),
strip.background=element_rect(color="white",
linewidth=0.5, linetype="solid"))
options(repr.plot.width=5.5, repr.plot.height=5)
print(Fig2)
ggsave("Fig2.png", plot= Fig2, width=5.5, height= 5, dpi= 300)

We aim to develop open-source code for agronomy ([email protected])
© 2022 – 2025 https://agronomy4future.com – All Rights Reserved.
Last Updated: 11/07/2023